Cloud solution
Don't worry if your machine has low resources.
All the software tools are executed in our server.
Multiple neuroscience fields
Always improving
We are always actively improving and developing new tools for NeuroSuites.
Feel free to contact us for help or suggestions.
Latest updates
In our last update we included multi-label classification capacity to supervised classification section to train, predict and compare multiple machine learning models for every kind of data set.
In our previous update we expanded our neuroscience fields to provide new tools for learning Gene Regulatory Networks (GRNs)
We are proud to introduce you our new set of machine learning tools including:
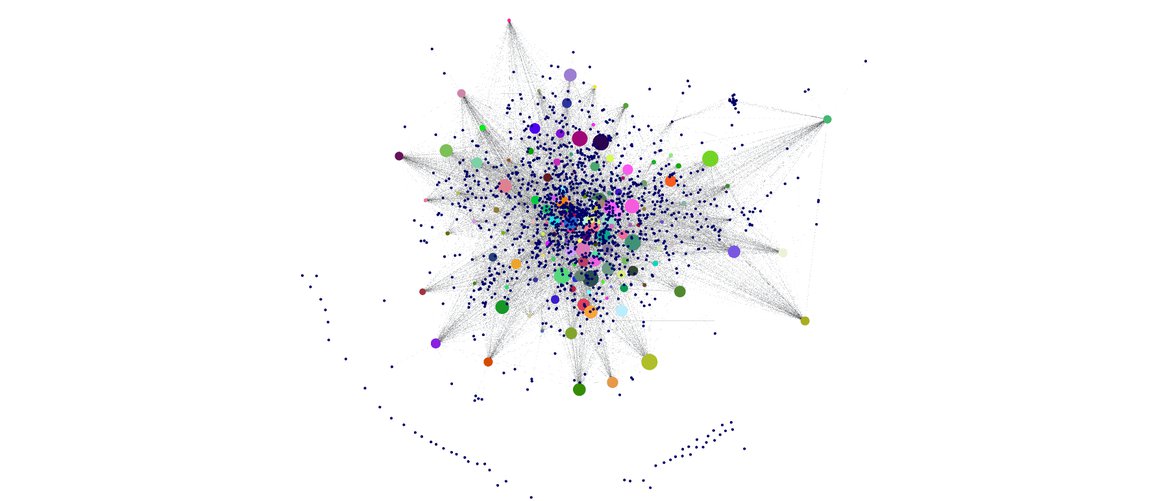
Bayesian Networks: structures and parameters learning and full visualization and inference
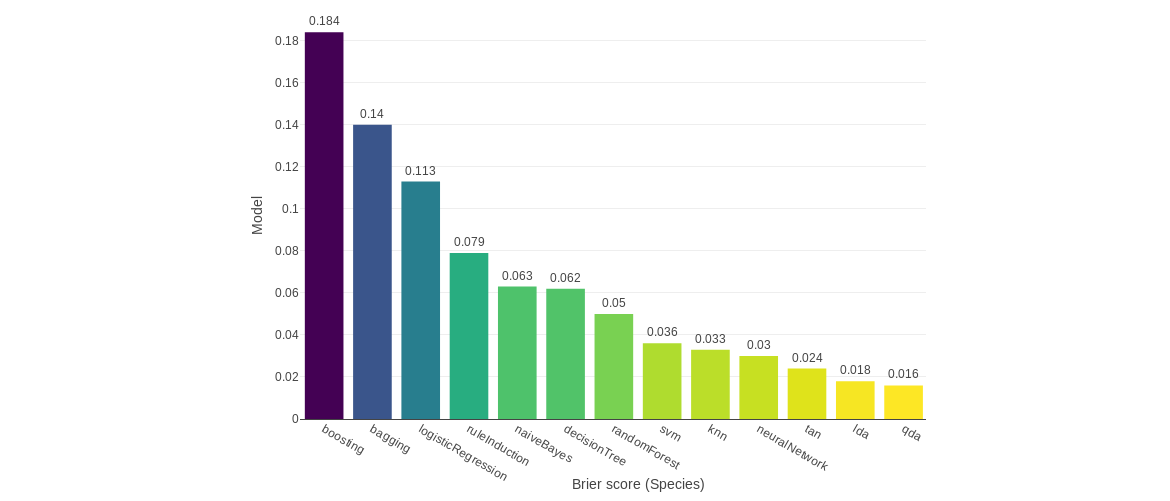
Supervised classification: data pre-processing, learning ,inference and evaluation
Probabilistic clustering: visualization and inference
Non Probabilistic clustering: visualization and inference
Some of these methods like our FGES-Merge have been specifically designed to infer the structure and parameters of large scales networks like GRNs. Some other methods are not bonded to specific application fields. We also included a large set of methods and tools to work with probabilistic graphical models in a general way, not only for GRN.
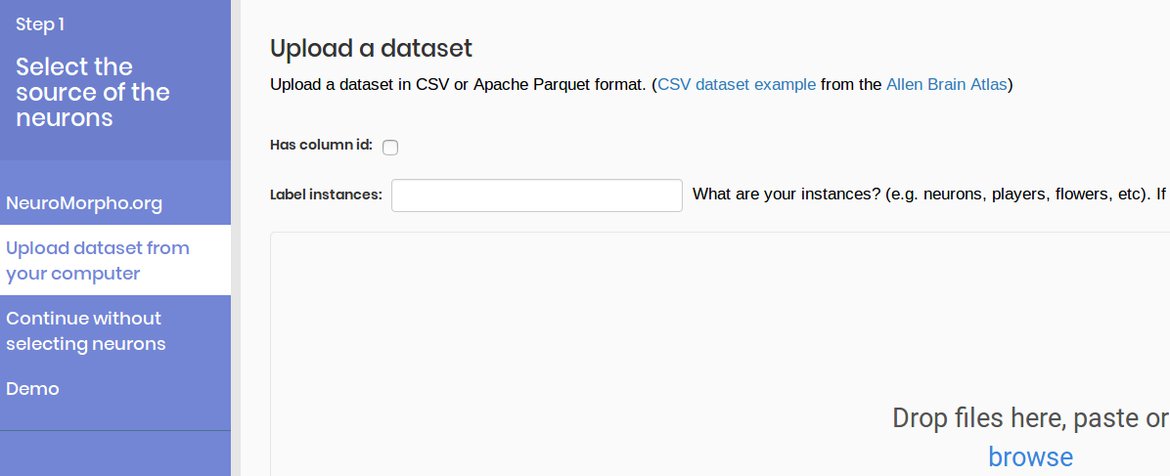
1. Upload your data set
You can upload upload your data set of your preferred scientific field.
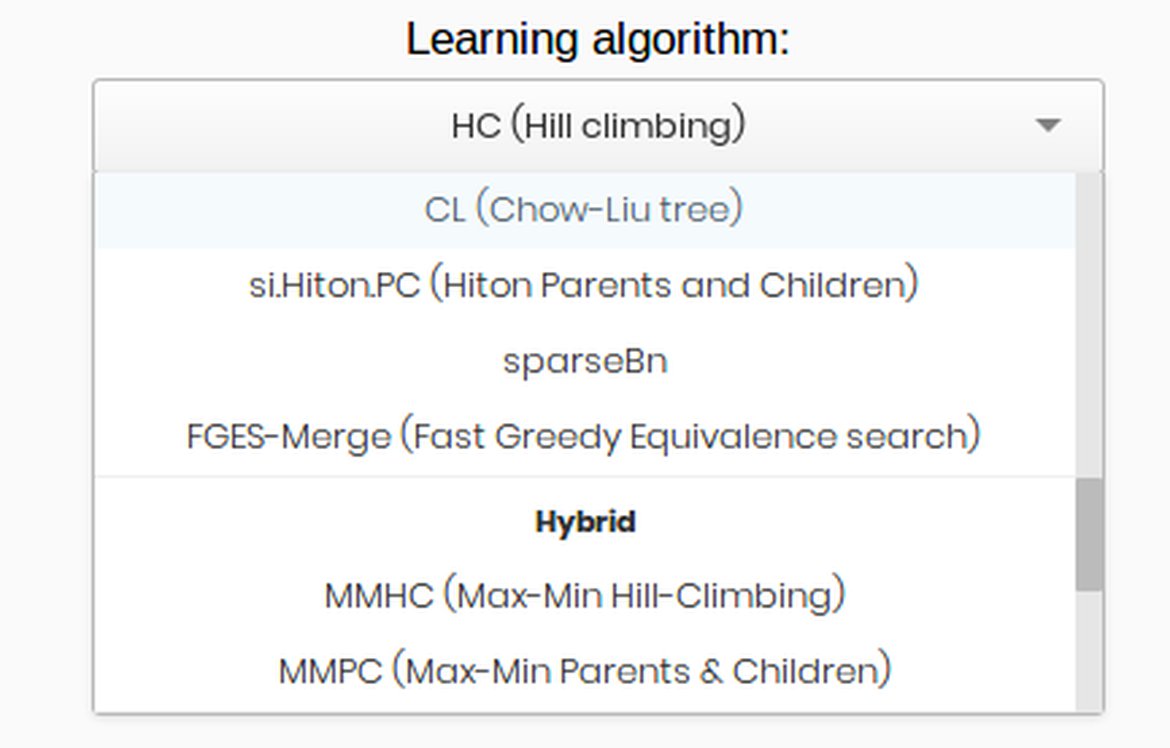
2. Learn the structure and parameters
Once the data set have been uploaded, go to the "Machine Learning" tab at the left side and click on "Bayesian Networks".
Select the desired features of your data set and click "Continue".
Then select the desired algorithm to learn the model structure. Once the structure has been learned you can also learn the continuous parameters of the model.

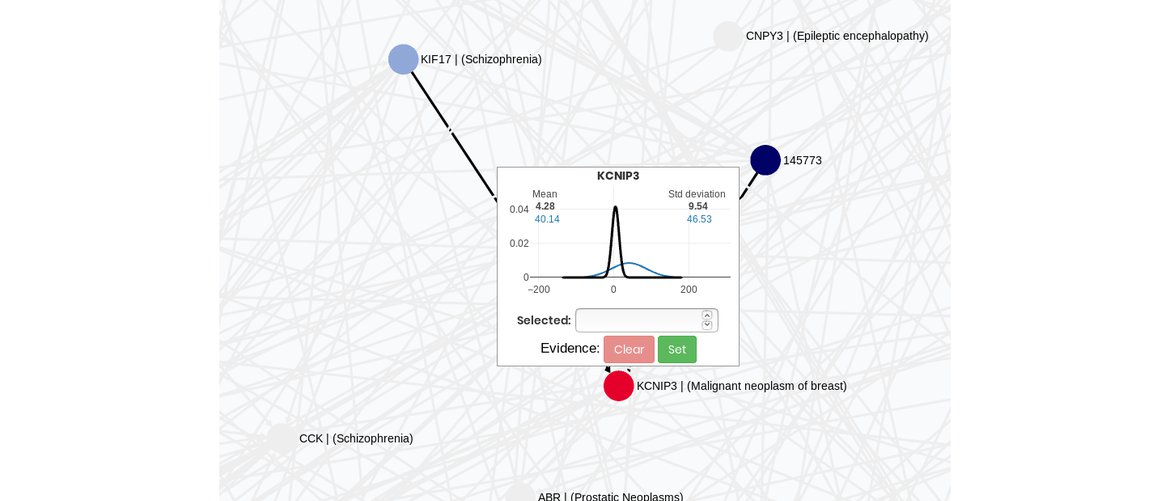
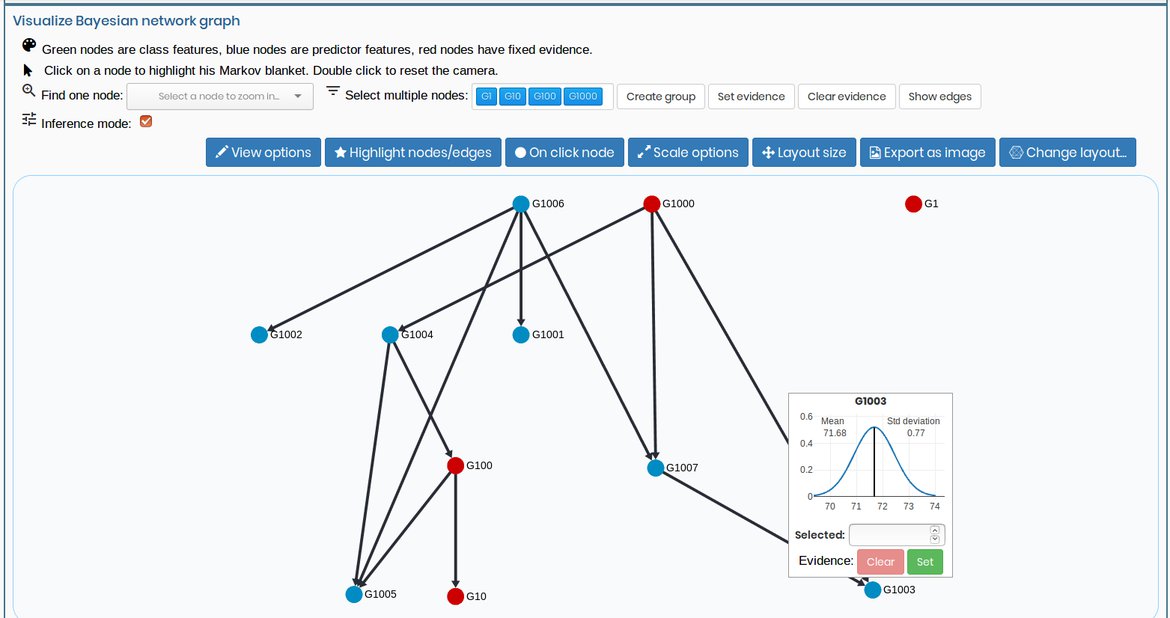
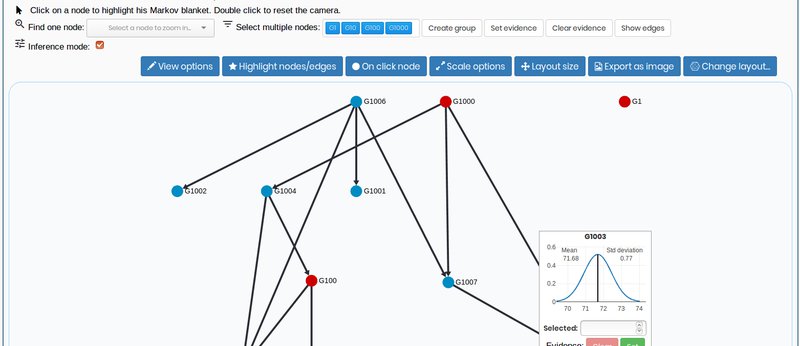
3. Visualize the model and make inference
View your probabilist graphical model in a full dynamic environment where you can also make inferences.
You can also upload your model already created externally to visualize it in our application.
What is NeuroSuites?
NeuroSuites is an online platform to run multiple neuroscience tools in a very easy way.
To start using NeuroSuites just click on your preferred category on the tab at the top of the page.
It provides you multiple tools to analyze neuroscience data. You will not need to install anything to run the tools provided here, everything is online. Furthermore, when you are done, you can export your results to your own computer.
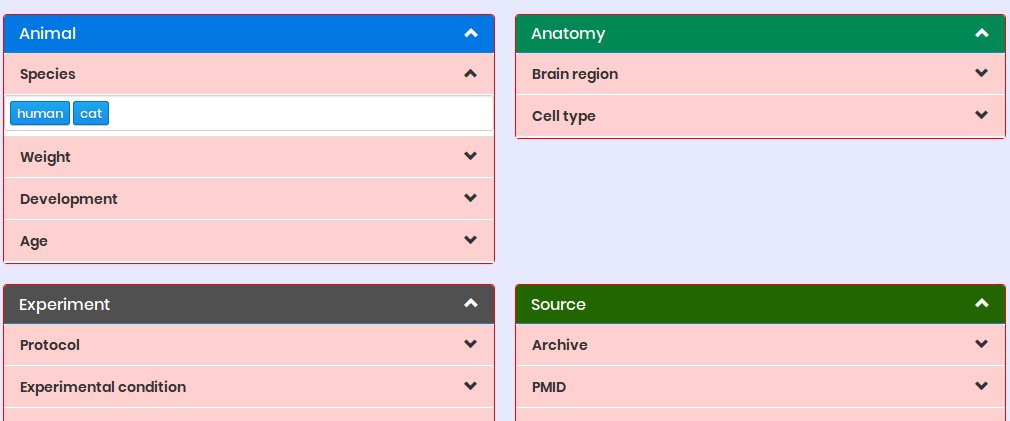
Who can use NeuroSuites?
Everyone can use NeuroSuites but it is intended to neuroscientists or data scientists that need to use neuroscience software in a fast and easy way without installing a lot of software in their computers.
No computer science or programming knowledge are required.
What services are available?
Machine learning
We have many available tools , some are focused in analyzing neurons morphology reconstructions and the others are general purpose tools like the statistics engine, supervised classification models, Bayesian networks, etc.
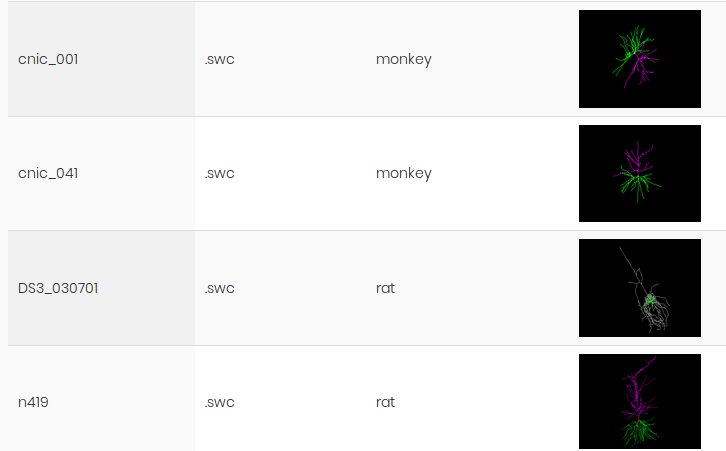
You can upload your own data set or you can select the neurons in the NeuroMorpho.org database.
Then you can use the following tools to analyze your neurons:
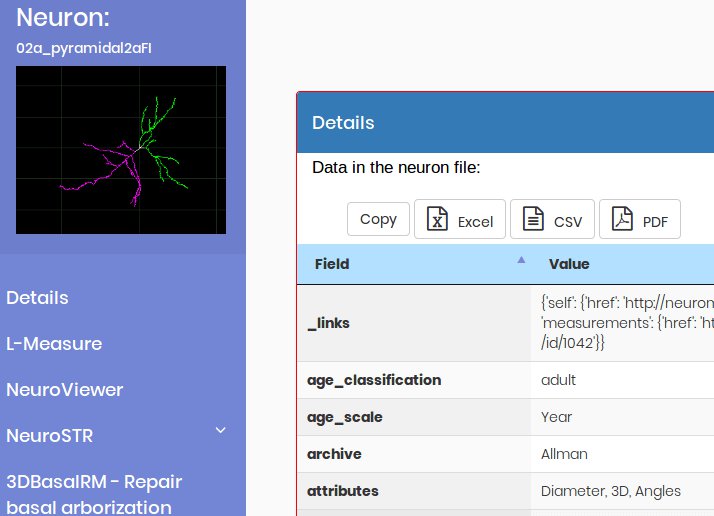
L-Measure - Extract morphological measurements
This tool allows researchers to extract quantitative morphological measurements from neuronal reconstructions.
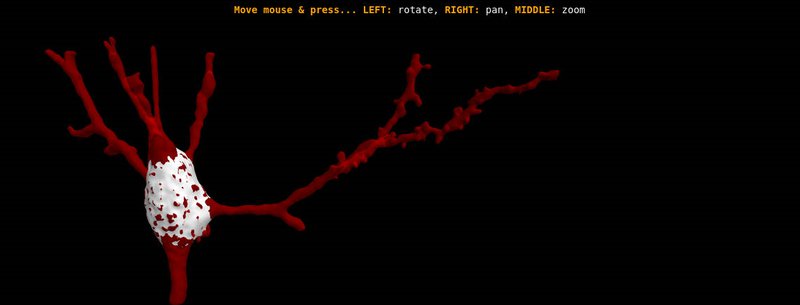
NeuroViewer - 3D Neuron reconstruction
3D Neuron reconstruction visualization package
Statistics engine
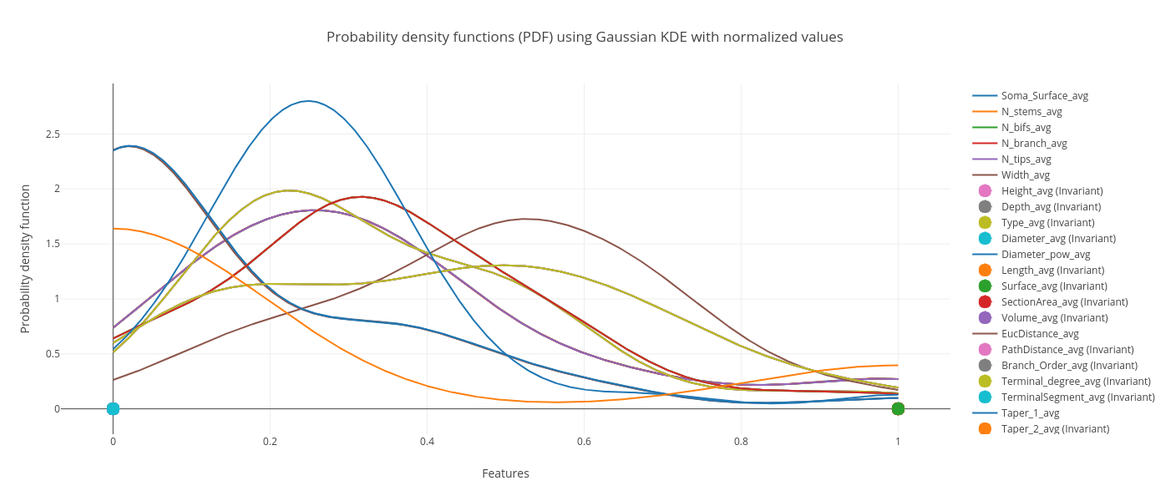
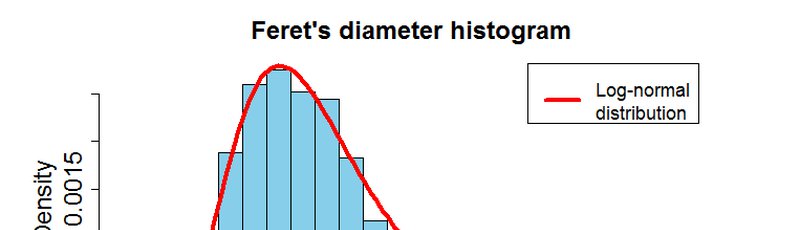
Descriptive statistics:univariate, bivariate and multivariate analysis and visualization.
Inferential statistics: confidence intervals, hypothesis testing (one sample t-test, two independent samples t-test).
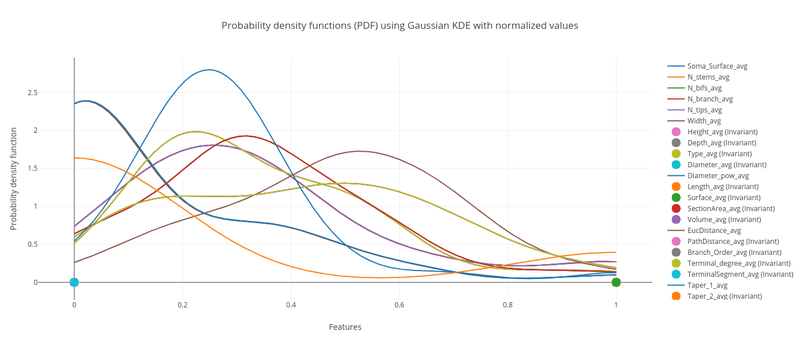
Interactive plots with Plotly (histograms, probability density functions, box plots, 2D and 3D scatter plots, Chernoff faces, Radar charts, Parallel coordinates, Andrew curves and much more!), custom options, exporting formats, etc. Everything online. Check out the L-Measure tool in the Morphometric Analyzer to see it in action.
Machine learning models
Bayesian Networks: structure and parameters learning for continuous and discrete datasets. Full visualization and inference for continuous BNs.
Probabilistic clustering graphical models: full visualization and inference for continuous models.
Supervised classification: learning and inference of multiple supervised classification models: k-Nearest neighbors, rule induction (CN2), decision tree, random forest, SVM, neural network, LDA, QDA, logistic regression, Naive Bayes, TAN, bagging meta-classifier, boosting (AdaBoost) meta-classifier, stacking meta-classifier. It also lets to perform classification for a multi-label classification problem using problem transformation approaches like binary relevance, classifier chains, label powerset, or RAKELd. The other approach for this kind of problems is with algorithm adaptation methods like multi-label knn or multi-label SVM.
Non probabilistic clustering: Discover groups in the data using multiple non probabilisitc clustering methods models: hierarchical agglomerative clustering, k-means, dbscan, affinity propagation and spectral. after it you can download the data with clusters and visialize them on a PCA with 2 components plot.
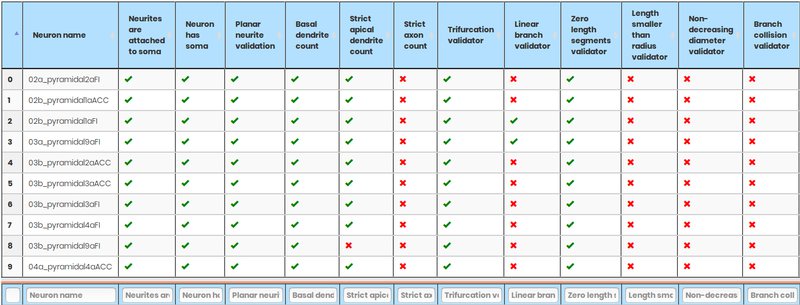
NeuroSTR - Validator, format converter
NeuroSTR is a neuroanatomy toolbox. It reads and processes three-dimensional neuron reconstructions in the most common file formats and offers a huge set of functions and utilities to work with them.
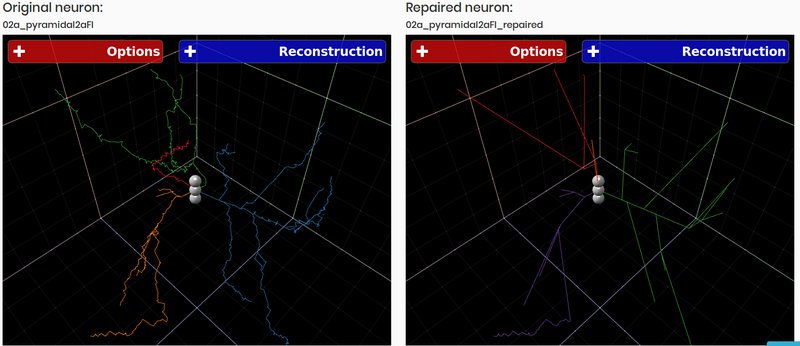
3DBasalRM - Repair cut-points in the basal arborization
Data-driven repairing model that detects cut-points in the basal arborization and then repairs them using a growth model built from complete three-dimensional neuron reconstructions As result a neuron in JSON format is returned.
GabaClassifier - Interneuron classifier
Classifies the given interneuron morphology into one of the 8 possible classes.
3DspineS - Dendritic spine simulation
This mathematical approach could provide a useful tool for theoretical predictions on the functional
features of human pyramidal neurons based on the morphology of dendritic spines.
3DSomaMS - Delimit the neuronal soma
This software provides a mathematical definition and an automatic segmentation method to delimit theneuronal soma.
3DSynapsesSA - Analyze spatial distribution of cortical synapses
Process and analyze patterns in the three-dimensional spatialdistribution of cortical synapses.
Dendrite arborization simulation
Generation of synthetic neurons with soma and dendrites.

Microscopy images
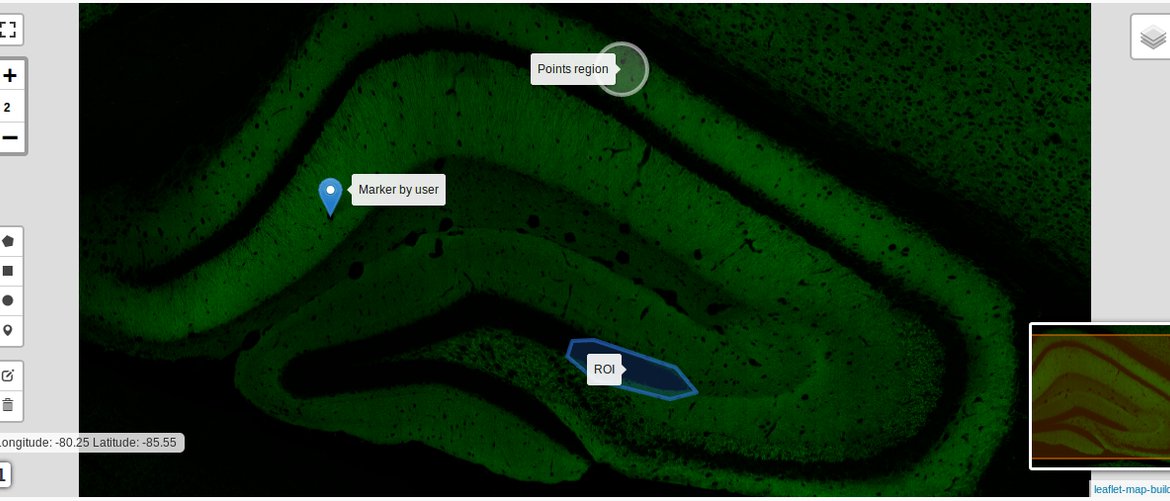
MultiMap
Multimap is an extensible application to create, visualize and analyze spatial data on maps.
Maps can be both geographical and non-geographical maps (for example maps created from biological images).
New tools and updates are coming...
Coming soon!
Release change logs
What is new in version 1.3 Released: 02/07/2020
What is new in version 1.2 Released: 03/06/2020
What is new in version 1.1 Released: 02/04/2020
What is new in version 1.0 Released: 12/11/2019
Version 0.9 Released: 07/23/2019
Version 0.8.1 Released: 02/22/2019
Version 0.8 Released: 10/15/2018
Version 0.7 Released: 08/25/2018
Version 0.6 Released: 07/27/2018
Version 0.5 Released: 07/06/2018
Version 0.4 Released: 06/29/2018
Version 0.3.1 Released: 06/19/2018
Version 0.3 Released: 05/31/2018
Version 0.2 Private release: 05/10/2018